Researchers developed a high-resolution spatial proteomics method to map protein and sugar changes in the brain. The study identified molecular signatures that distinguish healthy aging from Alzheimer’s and Lewy body disease.


In this really interesting essay, Michalon et al discuss defining Alzheimer’s disease in response to recent discussions on revising the definition and diagnostic criteria for the condition. The essay provides interesting historical context to the debate.
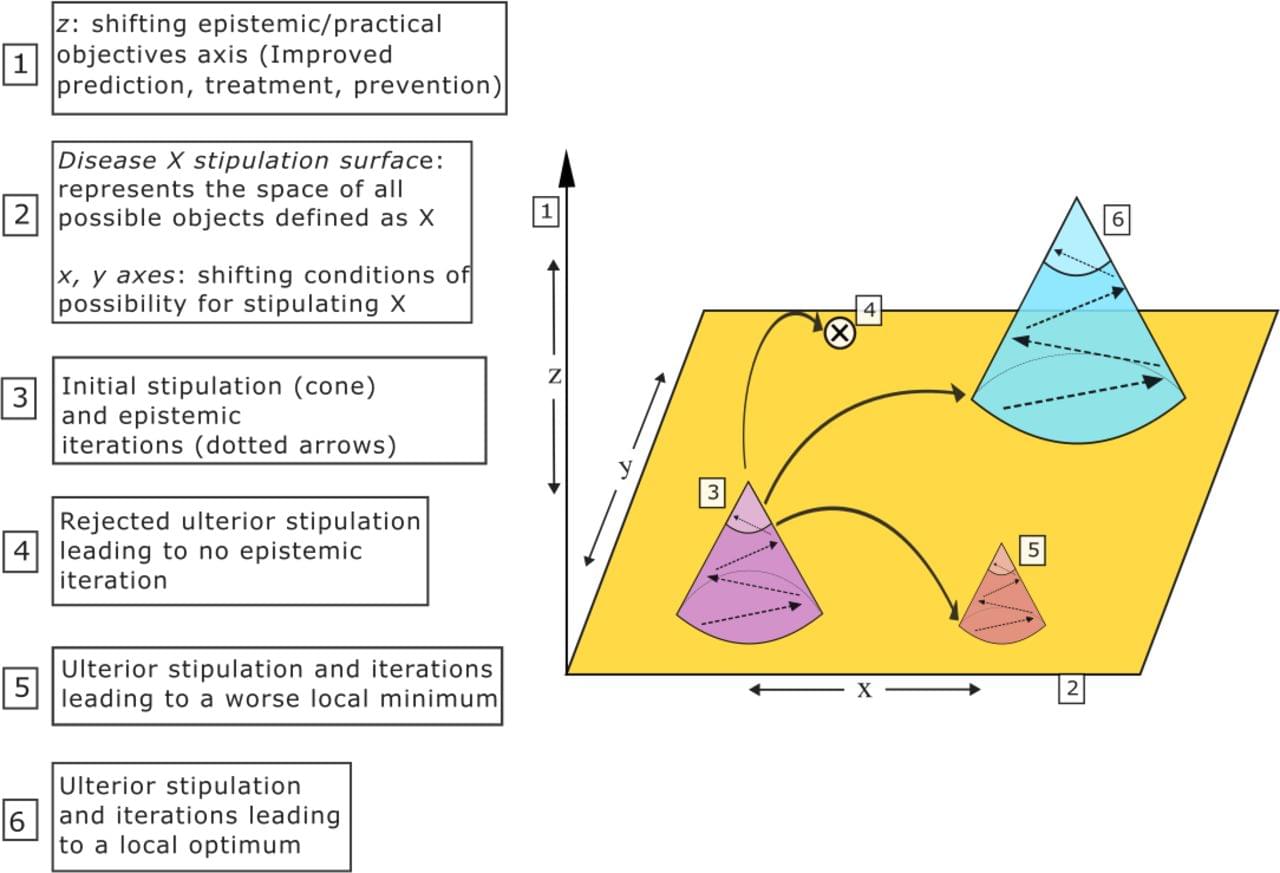
Recent revisions of Alzheimer’s Disease (AD) definitions by two leading research groups—the Alzheimer’s Association and the International Working Group—reflect divergent approaches: the former promotes a strictly biological definition, while the latter promotes a clinicalbiological construct. We contend that this emerging controversy is not merely semantic, but scientifically, clinically, and politically significant. Drawing on philosophical tools and situating the current debate within a broader historical context from the reconceptualization of AD in the 1970s onwards, we explore how definitions can serve as transformative instruments, acting as strategic bets that reshape scientific fields and clinical practices. Ultimately, we draw from the AD case study to argue for a critical reflection on the risks and promises of such definitional acts. We also propose a renewed attention to the ‘ethics of stipulating’ in the field of contemporary biomedical sciences.
In response to advances in diagnostics and therapeutics, two major research groups specialising in Alzheimer’s disease (AD) have recently revised their definition and diagnostic criteria for the condition. While they concur on certain aspects—most notably, the centrality of amyloid and tau pathologies—the two groups have proposed different types of definition. The Alzheimer’s Association (AA) group asserts the following fundamental principle: “AD is defined by its unique neuropathologic findings; therefore, detection of AD neuropathologic change by biomarkers is equivalent to diagnosing the disease” 1(p.5145). This definition regards specific biological changes as the unique defining feature rather than a joint characteristic, together with specific symptoms, of a disease. In this framework, asymptomatic individuals can be diagnosed with ‘preclinical AD’
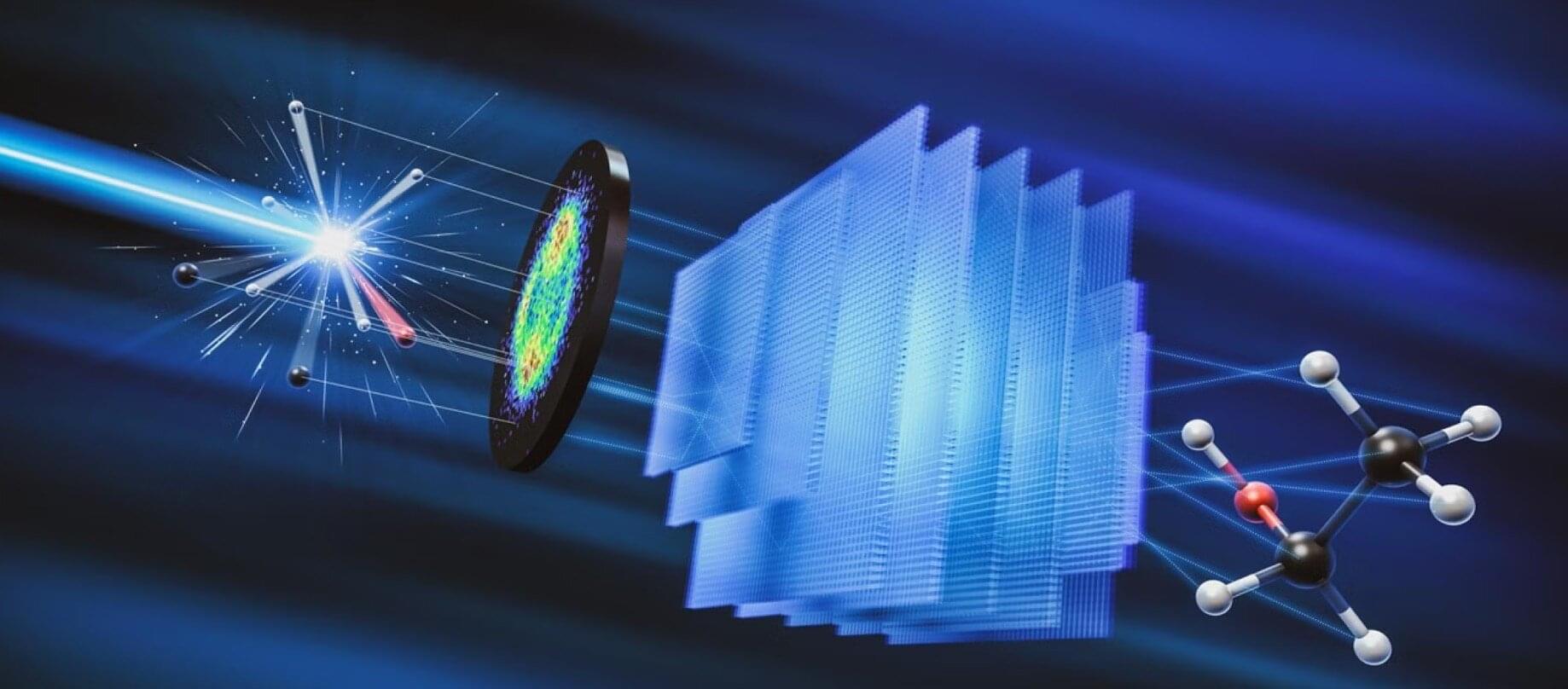

Researchers at the Department of Energy’s SLAC National Accelerator Laboratory and collaborating institutions recently built a generative AI model that can recreate molecular structures from the movement of the molecule’s ions after they are blasted apart by X-rays, a technique called Coulomb explosion imaging.
The research, published in Nature Communications, is an important step toward being able to take snapshots of molecules during chemical reactions—an advance that could have important impacts in medicine and industry. The machine learning model closely predicted the geometries of a range of different molecules made of less than ten atoms, paving the way for applying the technique to larger molecules.
“We were pretty excited about this,” said Xiang Li, an associate scientist at SLAC’s Linac Coherent Light Source (LCLS) and lead author of the study. “It is the first AI model built for molecular structure reconstruction from Coulomb explosion imaging.”
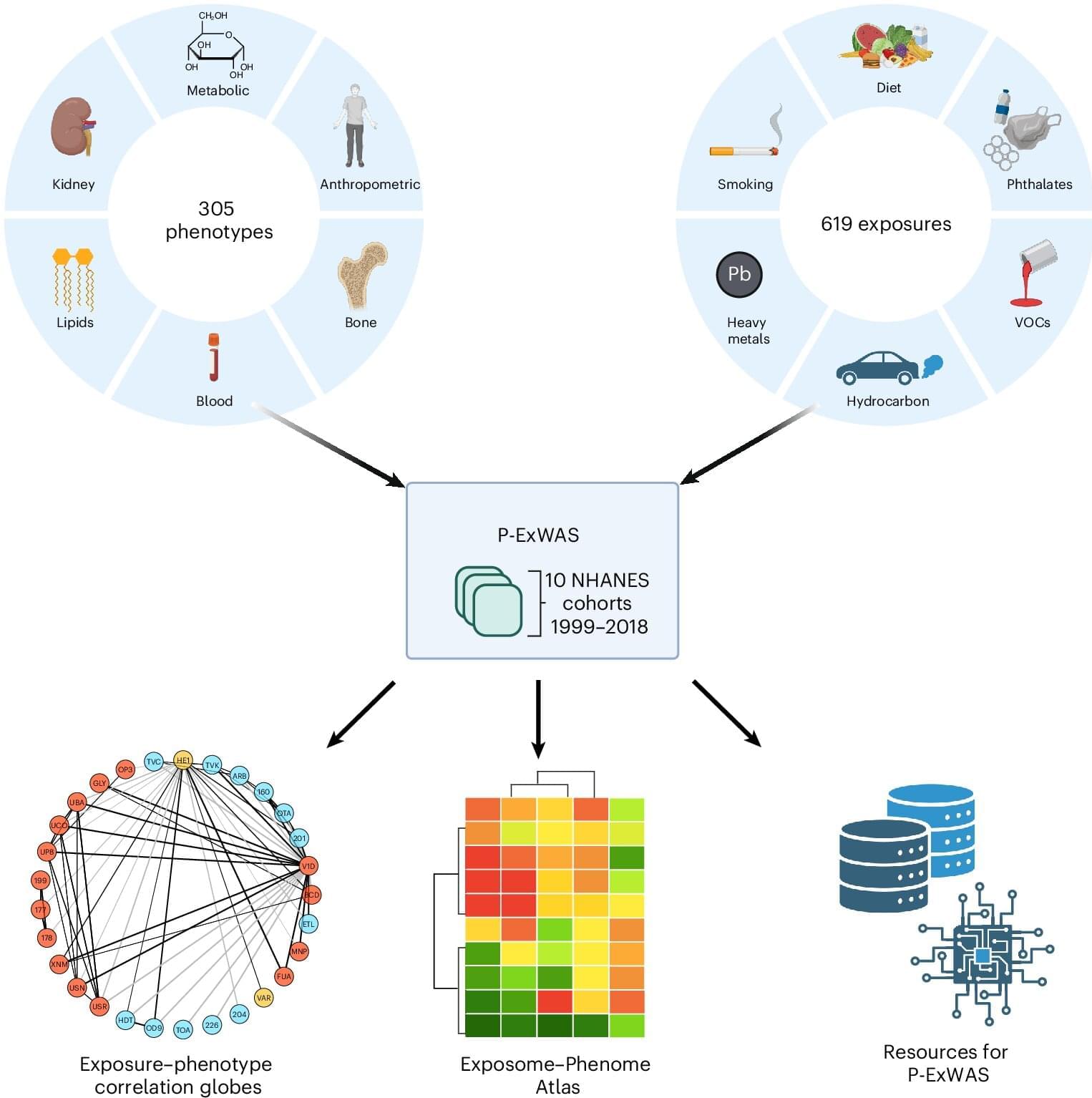
For decades, scientists have been carefully unraveling the role of genes in disease by examining how small variations in a person’s genetic code can shape lifelong risk of developing common conditions such as cancer, diabetes, or heart disease. But genetics only tell part of the story.
The other part comes from all the external and internal exposures a person experiences during their lifetime, which can range from pollution to infections to diet and lifestyle. Cumulatively, these exposures—and the body’s biological response to them—make up what scientists have termed the exposome.
A team led by scientists at Harvard Medical School has now conducted what may be the largest-scale study to date to quantify the relationships between exposures and health outcomes, testing more than 100,000 associations. The work demonstrates the importance of studying potential environmental disease risks in aggregate rather than one at a time.

Mass spectrometry is already a powerful tool for determining what kind and how many molecules are present in a given sample. But most instruments still analyze their molecules one or just a few at a time, an approach that is inefficient and costly, and in which rare, but significant molecules can easily fall between the cracks.
A more powerful version of the technology could one day allow scientists to read the full molecular contents of a single cell, track thousands of chemical reactions at once, and ultimately accelerate efforts like drug development.
Now, a new study describes the first big step in that direction by producing a prototype, dubbed MultiQ-IT, that’s capable of handling vast numbers of molecules at once. The findings, published in the journal Science Advances, offer a blueprint for faster, more sensitive instruments that could position mass spectrometry for the kind of transformation that reshaped genomics and computing.



Danièle Stalder, Conceição Pereira, David C. Gershlick et al. (University of Cambridge) uncover new regulators of Golgi-to-plasma membrane transport, showing that a PTPN23-dependent ESCRT pathway is essential for secretion of membrane & soluble cargoes, including hormones & antibodies.
Stalder and Pereira et al. combine affinity isolation of post-Golgi carriers, mass spectrometry, and a pooled CRISPR-KO screen to uncover new regulators of.
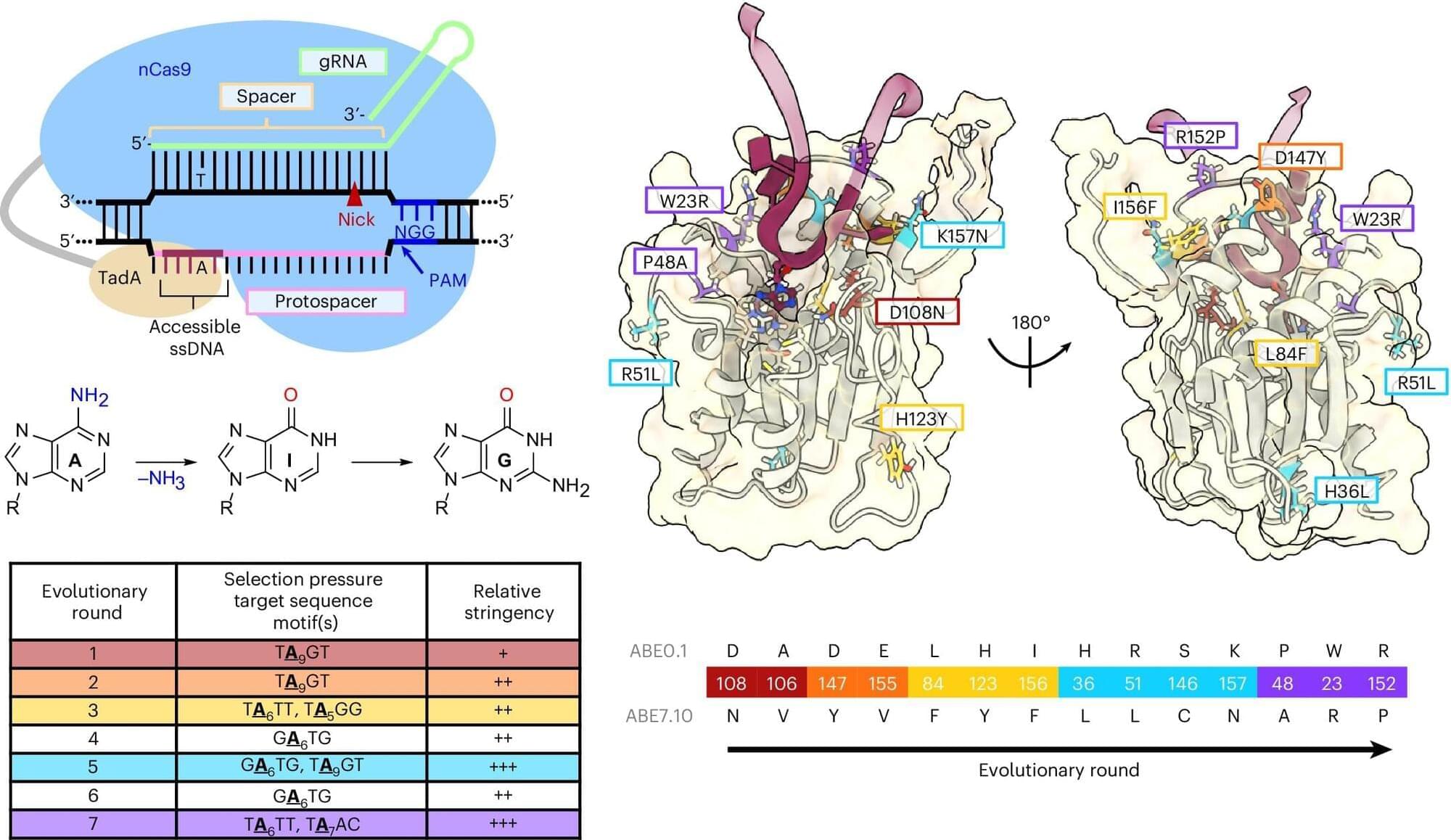
The trajectory of base editing has been remarkable, progressing from the laboratory to patient care, treating debilitating or terminal illnesses, in less than a decade. A type of gene editing that makes chemical changes to our DNA, base editing was developed by Alexis Komor, associate professor in the Department of Biochemistry and Molecular Biophysics at the University of California San Diego.
For all of base editing’s success, it is still a relatively new technology, and researchers like Komor are working to improve its efficiency, while lowering the incidence of unwanted edits. One type of unwanted edit is called a bystander edit. This occurs when a base editor not only edits the desired nucleobase, but also edits surrounding bases as well. Komor’s lab has developed a way to minimize bystander edits. This work appears in Nature Biotechnology.
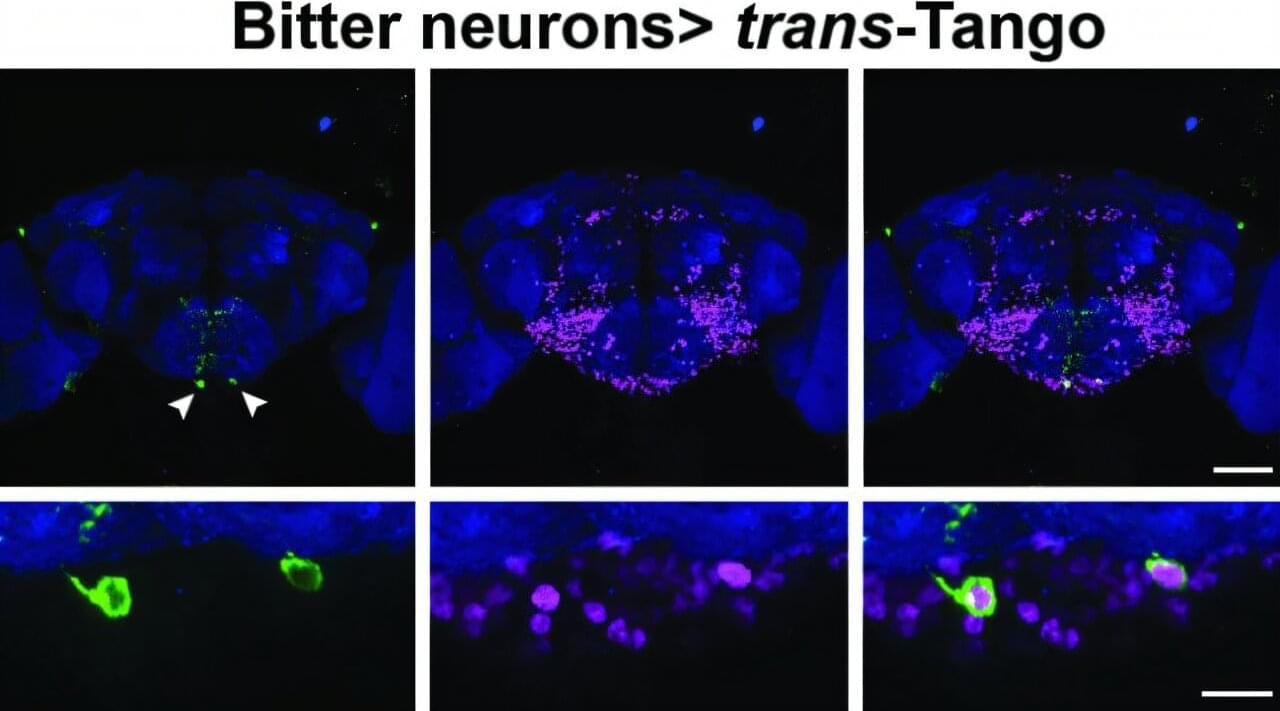
For the fruit fly, a sense of taste is critical to whether it thrives or dies. The little winged creature has taste organs in its mouthpiece as well as throughout its body, including its legs, abdomen and wing margins. When a fruit fly lands on a ripe or rotting fruit, it instantly receives information about whether the fruit is bitter or sweet. Sweetness indicates a caloric payday that cues the fly to feed; bitterness prompts the fly to move on from the potentially toxic substance.
Researchers in the lab of Brown University professor Gilad Barnea have identified the pair of neurons that make this critical choice. The insights on how flies navigate this complex decision-making process, a process not previously clear to scientists, are published online in Nature Communications.
“If a fly makes just one mistake about what to eat, it may die,” said Barnea, a professor of neuroscience and director of the Center for the Neurobiology of Cells and Circuits at Brown’s Carney Institute for Brain Science. “So the decision is super important. This newly discovered mechanism illustrates the impressive level of computation that a single neuron can do.”