Scientists uncover a new “recipe” that shows how exhausted T cells can be reprogrammed to regain their ability to attack tumors.


Chiari malformation (CM) involves a broad disease spectrum, where rare complex CM cases can be associated with craniocervical junction (CVJ) instability and require occipitocervical fusion. However, the natural progression of CVJ alignment in the general CM type I and 1.5 populations treated with posterior fossa decompression (PFD) remains insufficiently characterized. The authors aimed to compare CVJ alignment changes in patients who underwent PFD versus patients with CM who did not undergo surgery.
The authors conducted a retrospective cohort study at their institution of all patients diagnosed with CM I and 1.5 from 2000 to early 2023. Demographic, clinical, and surgical data were collected, along with preoperative and postoperative MRI measurements, including tonsillar herniation, brainstem descent, clivoaxial angle (CXA), and condylar-C2 sagittal vertical alignment (C-C2SVA).
A total of 241 patients were included, with 201 undergoing PFD and 40 managed conservatively (controls). No significant differences were observed between groups in age at diagnosis, sex, or genetic diagnoses. In the PFD group, 55% underwent duraplasty and 45% underwent bone-only decompression. Baseline craniocervical alignment measurements showed a lower CXA in the PFD group (144.4° ± 13.4°) compared to controls (148.5° ± 14.2°) (p = 0.04) but no difference in C-C2SVA. Changes over time showed a small but significant decrease in CXA at < 1 year after surgery in the PFD group (−2.7°) compared to controls (−2.0°) (p = 0.008), but no differences were noted at 1–2 years. No differences in C-C2SVA were observed over time in either group.
Aging doesn’t rewrite your DNA, it scrambles how your cells read it.
This clip explains epigenetic drift and how Life Biosciences’ therapy, ER100, uses Yamanaka factors to restore youthful epigenetic patterns in aged cells. By resetting the chemical marks that control gene expression, cells can behave as if they’re young again without changing the underlying DNA.
It’s the same biological process that happens early in embryonic development, applied in a controlled way to adult cells.

For life to develop on a planet, certain chemical elements are needed in sufficient quantities. Phosphorus and nitrogen are essential. Phosphorus is vital for the formation of DNA and RNA, which store and transmit genetic information, and for the energy balance of cells. Nitrogen is an essential component of proteins, which are needed for the formation, structure, and function of cells. Without these two elements, no life can develop out of lifeless matter.
A study led by Craig Walton, postdoc at the Center for Origin and Prevalence of Life at ETH Zurich, and ETH professor Maria Schönbächler has now shown that there must be sufficient phosphorus and nitrogen present when a planet’s core is formed. The study is published in Nature Astronomy.
“During the formation of a planet’s core, there needs to be exactly the right amount of oxygen present so that phosphorus and nitrogen can remain on the surface of the planet,” explains Walton, lead author of the study. This was exactly the case with Earth around 4.6 billion years ago—a stroke of chemical good fortune in the universe. This finding may affect how scientists search for life elsewhere in the universe.
Hopes are high for a powerful new compound aimed at lowering blood fat levels responsible for potentially fatal heart disease. In a recent trial, the oral drug, TLC-2716, lowered blood triglycerides by almost 40 percent and remnant cholesterol by more than 60 percent.
The drug was tested in a clinical trial involving 100 healthy adults to assess its effects on a metabolic switch that is active in the liver and gut, and involved in making and handling fats. The trial was the first of its kind on humans, and more testing is required.
Researchers initially isolated the switch, called Liver X Receptor ⍺ (LXR⍺), through analysis of large human genetics databases. Then they linked it to blood-fat-related metabolic disorders using Mendelian randomization, a powerful technique for linking gene expression and outcomes.
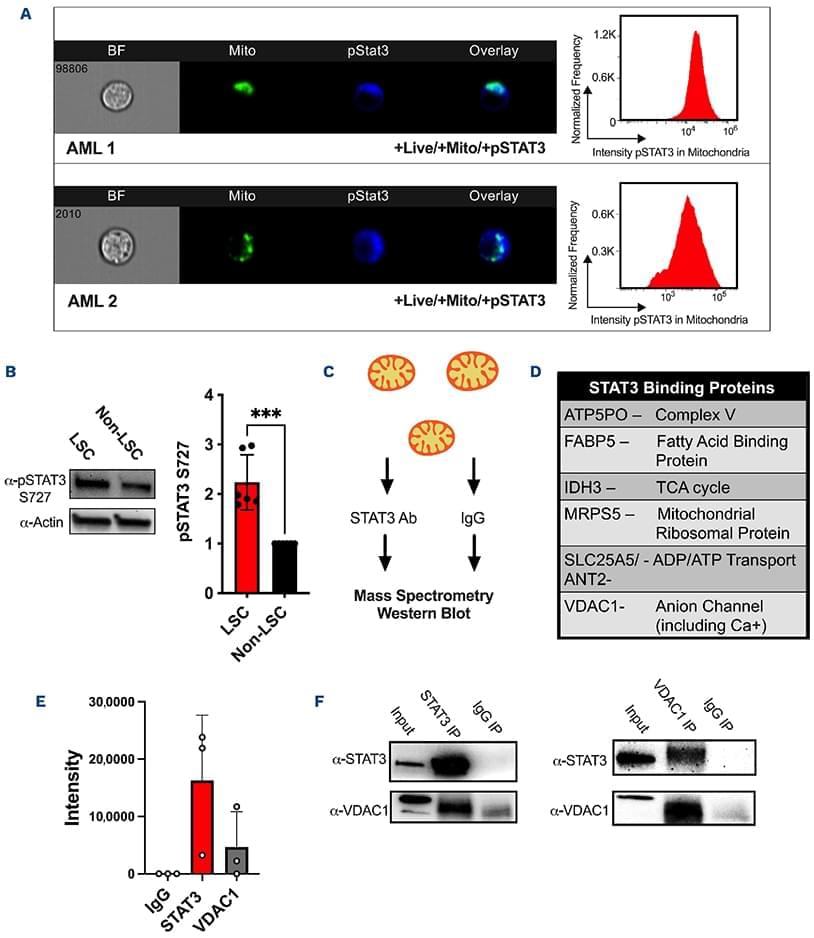
New insights into the role of signal transducer and activator of transcription (STAT)-3 in regulating mitochondrial function in acute myeloid leukemia (AML): by linking STAT3 signaling to mitochondrial metabolism and apoptosis control, this study provides new mechanistic insight into how AML cells maintain their energy balance and resist cell death. These findings highlight mitochondrial regulation as a potential therapeutic vulnerability in AML.
Signal transducer and activator of transcription 3 (STAT3) is a well-described transcription factor that mediates oxidative phosphorylation and glutamine uptake in bulk acute myeloid leukemia cells and leukemic stem cells. STAT3 has also been shown to translocate to the mitochondria in acute myeloid leukemia cells, and phosphorylation at the serine 727 (pSTAT3 S727) residue has been shown to be especially important for the mitochondrial functions of STAT3. We demonstrate that inhibition of STAT3 results in impaired mitochondrial function and decreased leukemia cell viability. We discovered a novel interaction of STAT3 with voltage-dependent anion channel 1 (VDAC1) in the mitochondria which provides a mechanism through which STAT3 modulates mitochondrial function and cell survival. Through VDAC1, STAT3 regulates calcium and oxidative phosphorylation in the mitochondria. STAT3 and VDAC1 inhibition also results in significantly reduced engraftment potential of leukemia stem cells, including primary samples resistant to venetoclax. These results implicate STAT3 as a therapeutic target in acute myeloid leukemia.
Acute myeloid leukemia (AML) is a genetically heterogenous and highly aggressive myeloid neoplasm with poor prognosis.1,2 Standard therapy for AML has historically consisted of induction chemotherapy with an anthracycline and cytarabine, followed by consolidation with either hematopoietic stem cell transplant or high-dose cytarabine.3 Recently, therapeutic options have broadened with the advent of novel targeted therapies.4–7 However, despite high response rates, relapse is common.6 Relapsed disease is believed to originate from a quiescent subpopulation of therapy-resistant leukemic stem cells (LSC)8 which are found in greater abundance at the time of relapse than at diagnosis,9–12 and negatively correlate with survival.10,11 LSC demonstrate a unique vulnerability in their preferential reliance on mitochondrial activity and oxidative phosphorylation (OXPHOS).12–14 While Bcl-2 inhibition with venetoclax in combination with the hypomethylating agent azacitidine has demonstrated selectivity for LSC through inhibition of OXPHOS,13 resistance frequently develops via alterations in mitochondrial metabolism or activation of alternative anti-apoptotic pathways.15–19 Furthermore, prior studies of patients who progress after frontline hypomethylating agent/venetoclax have shown very poor outcomes, with a median survival following failure of this combination of 3 months or less.20–22 New strategies targeting LSC via their reliance on OXPHOS are of significant interest and have been described in several reports,7,13,23 however, further research is needed to elucidate the mechanisms underlying the observations.
Signal transducer and activator of transcription 3 (STAT3) has been shown to be important for leukemogenesis and is known to be highly expressed in many AML patients’ samples and cell lines.24–27 Canonically, STAT3 is known to undergo phosphorylation at residue Tyr705 leading to dimerization and translocation to the nucleus where it functions as a transcription factor regulating cell development, renewal, proliferation, and cell death.25,28–30 Our previous work additionally established that the transcriptional activity of STAT3 regulates mitochondrial function via a MYC-SLC1A5-mediated pathway.27 Despite its well-described nuclear role as a transcription factor, STAT3 has also been discovered to localize to the mitochondria.31,32 Prior work has suggested a variety of functions in the mitochondria, including modulation of electron transport chain activity,31–33 regulation of mitochondrial genes,34 and regulation of mitochondrial calcium flux.35,36 While phosphorylation of STAT3 at both Tyr705 (pSTAT3 Y705) and Ser727 (pSTAT3 S727) sites have been found in the mitochondria,31–33,36,37 Ser727 phosphorylation is critical for modulation of mitochondrial functions such as electron transport chain activities.31,32 These data suggest that STAT3 plays a critical role in mitochondria, although this role in AML is not well characterized.
Join us on Patreon! https://www.patreon.com/MichaelLustgartenPhD
Discount Links/Affiliates:
Blood testing (where I get the majority of my labs): https://www.ultalabtests.com/partners/michaellustgarten.
Blood testing with LifeExtension.com: https://www.anrdoezrs.net/click-101614996-15750394
At-Home Metabolomics: https://www.iollo.com?ref=michael-lustgarten.
Use Code: CONQUERAGING At Checkout.
Clearly Filtered Water Filter: https://get.aspr.app/SHoPY
Epigenetic, Telomere Testing: https://trudiagnostic.com/?irclickid=U-s3Ii2r7xyIU-LSYLyQdQ6…M0&irgwc=1
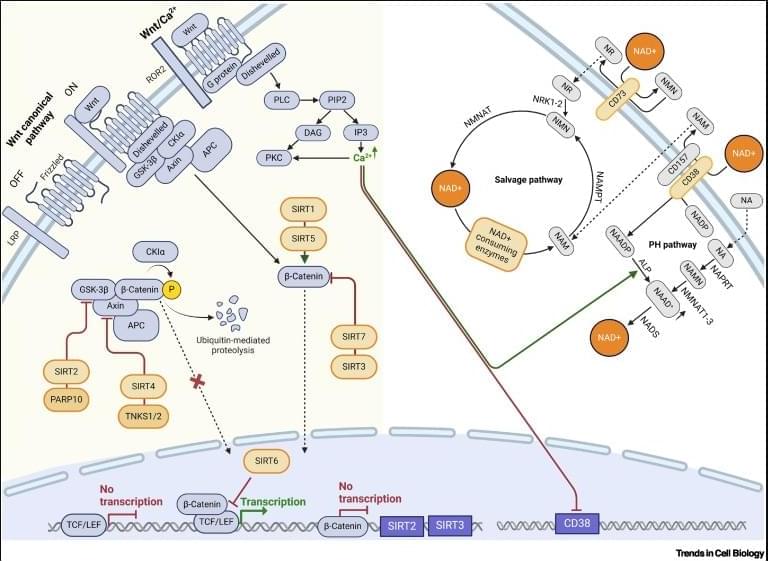
Wnt–NAD+ axis in stem cell function.
The Wnt–NAD+ axis is a fundamental regulatory hub in which metabolic state meets developmental signaling and it acts as a metabolic sensor that coordinates tissue regeneration with cellular energy status through compartment specific NAD+ pools.
Wnt signaling regulates NAD+ metabolism by controlling the expression of key biosynthetic enzymes and NAD+ consumers, while NAD+-dependent proteins modulate Wnt activity through direct interactions and epigenetic modifications.
Sirtuins exhibit tissue-specific and subcellular compartment-dependent roles in Wnt regulation where they function as activators or suppressors depending on the cellular bioenergetic state.
The Wnt–NAD+ axis maintains stem cell function and self-renewal capacity through metabolic/signaling integration, and its disruption during aging leads to declining regenerative capacity.
The progressive dysregulation of compartment-specific Wnt–NAD+ coordination contributes to stem cell exhaustion and multiple pathological conditions, indicating that therapeutic strategies must consider tissue-specific and subcellular targeting. sciencenewshighlights ScienceMission https://sciencemission.com/Wnt%E2%80%93NAD-axis
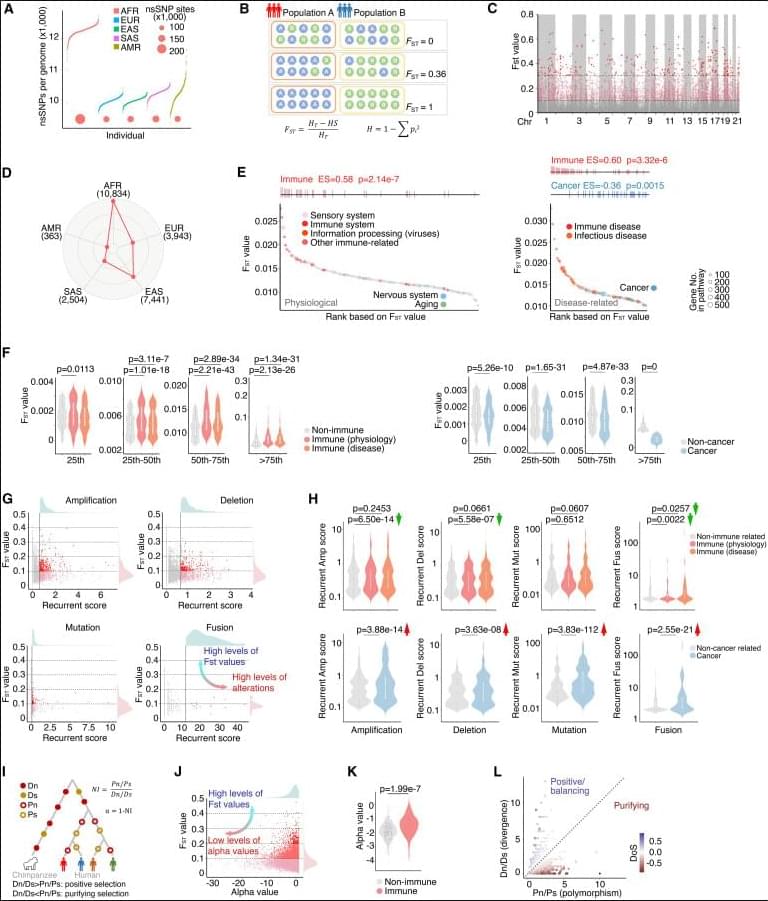
Hu et al. analyzed non-synonymous SNPs across diverse human populations and revealed divergent evolutionary pressures on immune-and cancer-related genes. By integrating population diversity with functional evaluation, they identified STING1 variants as modulators of interferon signaling. Their findings suggest that germline variations shaped by genetic ancestry may influence cancer immunity.
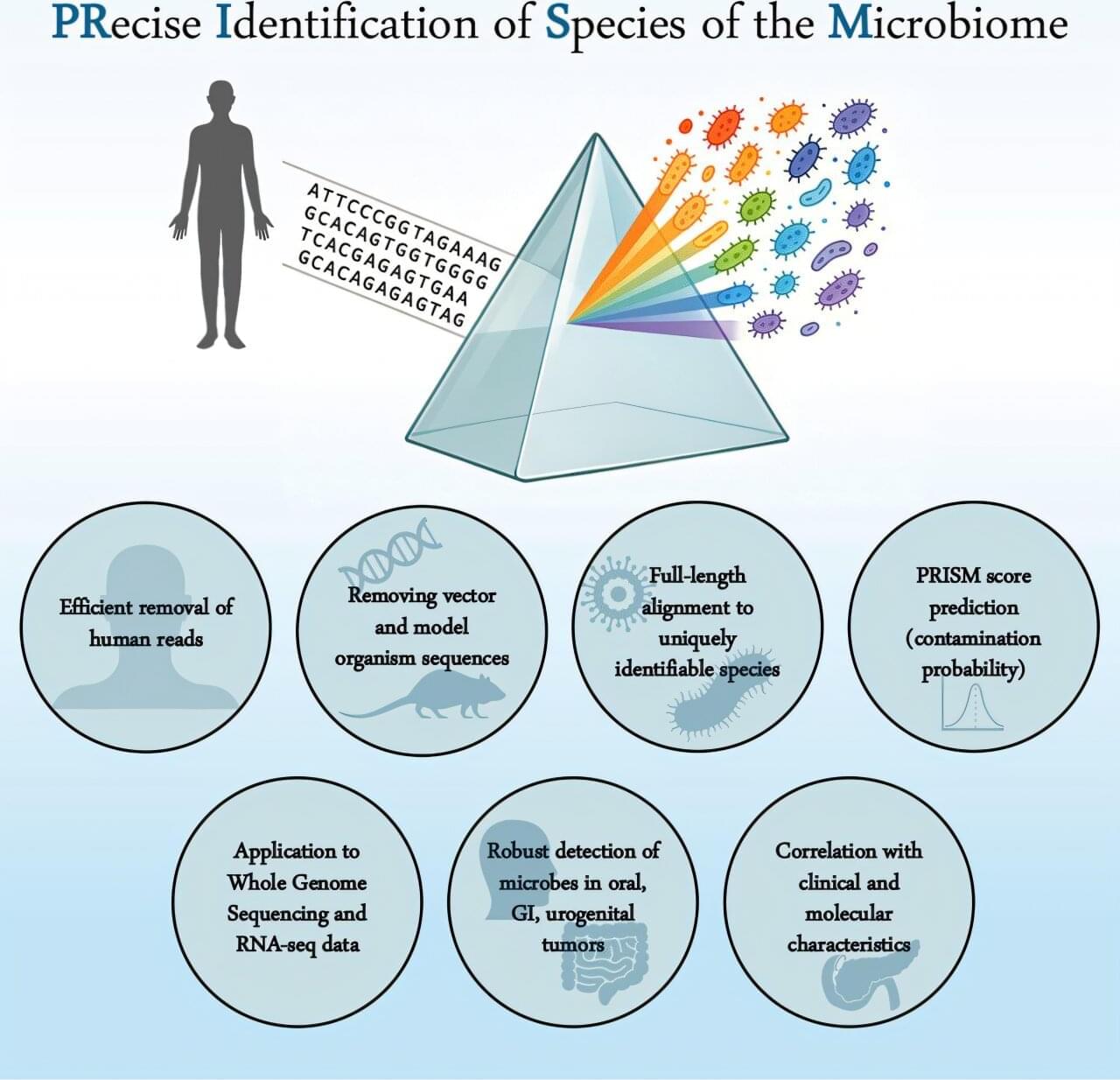
When scientists sequence tumor DNA, they typically find small amounts of genetic code from bacteria, viruses and fungi—microorganisms that—if actually present in tumor tissues—could influence how they grow, evade immunity or respond to treatment. But do microorganisms truly reside in tumors, or do the samples become contaminated before sequencing occurs?
Independent analyses of the same genomic data have reached wildly different conclusions. Now, researchers at Rutgers Cancer Institute have developed a computational tool that settles the controversy by distinguishing genuine microbial signals from artifacts. Their findings are published in Cancer Cell.
“There are microbes all over the environment, on our skin and in our breath,” said Subhajyoti De, a member of the Genomic Instability and Cancer Genetics Program at Rutgers Cancer Institute and the senior author of the study. “There could be DNA particles floating in the air. How do you know whether you’re finding came from the tissue you were interested in, or whether something was introduced along the way?”